Xuecai Ge is interviewed for the NIGMS Biomedical Beat Blog. Read to learn about Dr. Ge and the research performed by her lab. Congratulations Dr. Ge!

Stephanie Woo is a featured speaker at this year’s British Society for Developmental Biology Meeting in Sheffield UK. Awesome Stephanie!

New for 2023: UC Merced Developmental Biology and Neurobiology labs are holding joint lab meetings and journal clubs. Looking forward to learning something new and making new collaborations. Thanks Xuecai Ge for organizing!

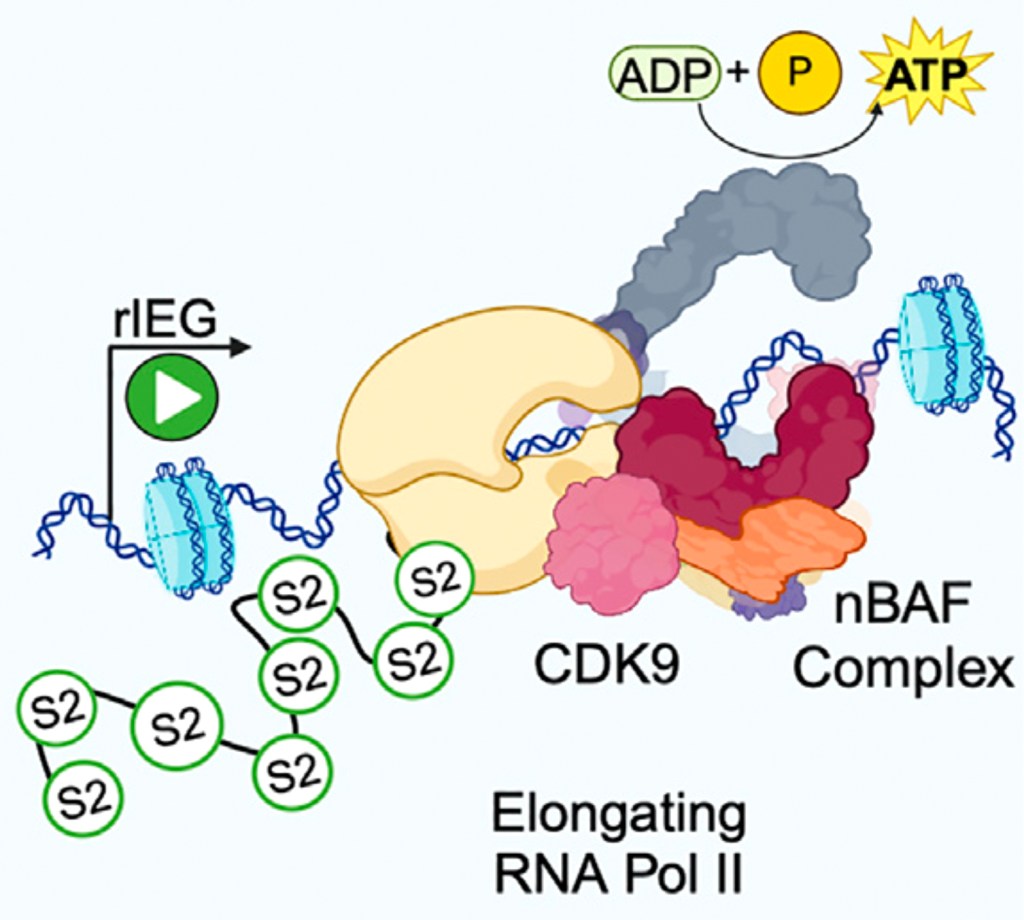
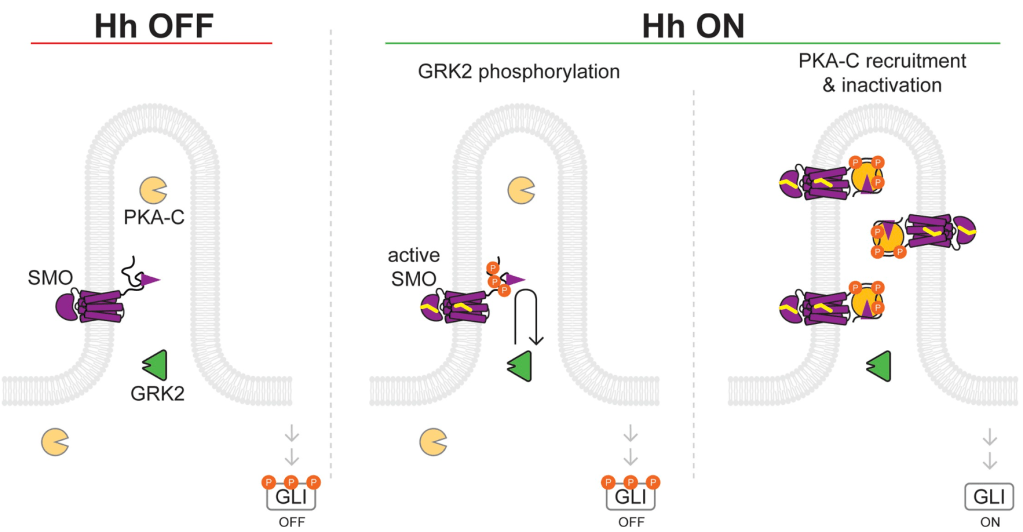
The UC Merced Neuro labs go to SFN! Ramen Saha (036.04/D6) Nov 12 1-5pm for nBAF and IEGs, Eva Cai (107.09/A8) Nov 13 8-noon for Hh signaling in cilia, Cal Larnerd (150.06/OO5) Nov 13 8-noon for EtOH tolerance memories, Xiaoliang Liu (606.24/C56) Nov 16 8-noon for Numb promoting Hh signaling, and Brian Wang (728.28/HH11) Nov 16 1-5pm for sex-specific thirsty water seeking pathways!

Fred Wolf was elected Chair of the Quantitative and Systems Biology Graduate Group at UC Merced. Learn more here.

Chris Amemiya is co-organizing the joint meeting of the Pan-American Society for Evolutionary Developmental Biology and the Society for Developmental Biology, in Vancouver BC, July 17-22, 2022. Abstracts are due April 11. Learn more here.